Codon: Notes on Progress #2
Nano-scale probes to explore the universe, gene-editing protects the heart, 217 new papers this month.
Notes on Progress is a monthly roundup of papers and ideas about biology and the future.
“The recombinant DNA breakthrough has provided us with a new and powerful approach to the questions that have intrigued and plagued man for centuries. I, for one, would not shrink from that challenge.”
— Paul Berg, Nobel Prize Lecture, 1980
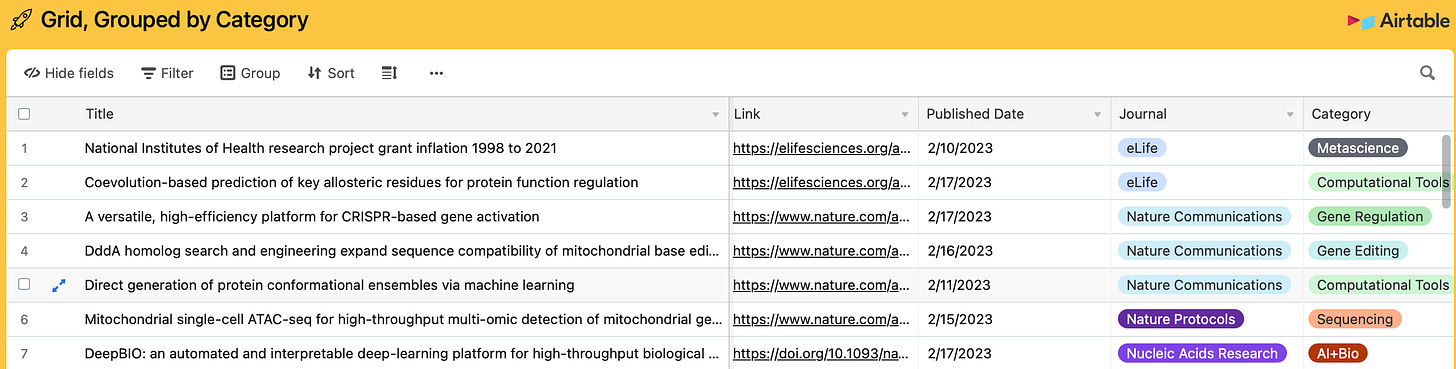
1 of 31. Every weekend, for the last four weekends, I’ve idled away my Saturdays at a computer keyboard looking through journals. My goal, ostensibly, is to find new papers about gene editing, neurotechnology, and AI and then add them to a public AirTable. I’ve added 217 studies to the database in the last 30 days. I envision this growing collection as a way to track scientific progress over time. I’ll update the list retroactively, too; I’ve covered thousands of scientific papers over the last three years of writing this newsletter.
(Tip: Click on “View larger version” in the bottom-right corner of the AirTable to view the database in full screen. Click on columns to filter by category, journal, or date.)
Since the last Notes on Progress, Codon has published two essays:
De-Extinction? Surely You’re Joking! A critical — but, in other ways, uplifting — view on “bringing back the woolly mammoth.” De-extincting a mammoth will not happen in the next ten years (I’d give it a 0.1 percent chance.) But the technologies devised in the process of trying may revolutionize other areas of science, especially reproductive biology.
Biology is a Burrito. Cells are crowded and very fast places. It's a miracle that anything works at all. The central dogma is often depicted as DNA→RNA→protein, but it’s much more: A biophysical marvel inside the smallest of vessels.
2 of 31. A recent study from George Church, the Harvard biologist (and de facto de-extinction leader), lays out what would be needed to make interstellar “probes” that could be launched into space and seed distant planets across the Universe with life. The paper takes a quantitative approach: How would these probes propel themselves? How much could they weigh? How would they decelerate and land on planets? How would they avoid crashing into interstellar dust? And how, upon arrival, would they communicate back to Earth?
Biology provides answers to some of these questions. Key takeaways:
It’s likely cheaper to launch one quadrillion probes that each weigh one picogram than to launch 1,000 devices that each weigh one gram.
A probe weighing one picogram could be propelled using a solar sail that measures just 10-8 square meters. For context, that area is 225,000 times smaller than a Post-it note.
Just 13 percent of probes would actually make it to a distant planet. Most probes would crash and burn due to collisions with dust and debris.
The probes could carry bacteria that emit flashes of light (via bioluminescence) upon arrival on a distant planet. Some bacteria can survive temperatures up to 420 degrees Celsius (specifically, spores of Bacillus amyloliquefaciens) or down to -20 degrees C.
Now who’s gonna build ‘em?
3 of 31. Two separate papers this month used a gene-editing tool, called base editing, to prevent hypertrophic cardiomyopathy in mice. In each case, animals carried a mutation in a gene that encodes β-myosin, a protein that helps heart muscles contract. Mutations in this gene cause hypertrophic cardiomyopathy — which affects 1 in 500 people — and thickening of the heart’s left ventricle. Hypertrophic cardiomyopathy is the leading cause of sudden cardiac death in people under 35 years.
Base editor proteins were delivered into the mice using a virus, called AAV9. The proteins fixed the mutation between 30 and 80 percent of the time. Mice had “complete rescue of heart wall thickness,” according to co-author David Liu, and “treated mice remained indistinguishable from [wildtype] mice throughout the 32-to 34-week study.” Nature Medicine (1 and 2)
4 of 31. I recently visited the MIT Museum, which has large exhibits devoted to biotechnology, automation, and artificial intelligence. The museum holds Claude Shannon’s original Endgame chess machine and also the first model of a tRNA molecule, which carries amino acids to ribosomes to make proteins. The tRNA model was built in 1975 by a first-year undergraduate student, using screws and pieces of metal.
I stood before this model, in awe, for at least ten minutes. I’ve also seen the Watson/Crick DNA model in London’s Science Museum, but I think this tRNA model is much better. Incredibly complex and delicate, it made me think about how far biology has come in just fifty years. Our greatest technologies to model biology went from scraps of aluminum to computers and AI tools that we already take for granted.
It’s good to be a biologist, but even better to be a biology writer, with the freedom to unceasingly search for new things under the sun.

5 of 31. Many new gene-editing tools published this month, including:
A more specific version of Cas9 (called Sniper2L).
A toolbox of seven (SEVEN!) gene-editing proteins that can be switched on- or off with a chemical. The kit includes two cytidine base editors, two adenine base editors, a prime editor, and a transcriptional activator.
Prime editing uses a modified form of the Cas9 protein to edit, insert, or delete portions of a DNA strand. It was used, in a recent study, to “fix” a genomic site using 3,604 different DNA insertion sequences. The experimental data was then used to train a machine learning model that could “predict relative frequency of insertions into a site with R = 0.70.” That’s quite good.
A new base editor to edit mitochondrial DNA.
A type of Cas protein, called Cas12j, can make allele-specific DNA edits (e.g. leave one copy of a gene alone, while “fixing” a mutation in the other copy). It could theoretically be used to correct “25,931 clinically relevant variants.”
Nearly every protein now used in genome editing (including Cas9, Cas12, and Cas13) came from bacteria. A new paper modifies an archaeal enzyme, called TnpB, to cleave DNA in much the same way as Cas proteins, but at very wide temperature ranges — between 37 and 85 degrees Celsius.
Related news: “More than 200 people have been treated with experimental CRISPR therapies.” The future of precision genetic medicine is far closer than many realize.
6 of 31. This newsletter is dedicated to the memory of Paul Berg, who died last month at the age of 96. Berg’s work set the stage for the entirety of synthetic biology and genetic engineering. His group was the first to make recombinant DNA, in 1971, by splicing E. coli genes into a piece of simian virus DNA. The landmark paper enabled revolutions in insulin biomanufacturing, CAR-T therapies, and the bioproduction of most modern vaccines; in other words, much of what we take for granted today. The New York Times

7 of 31. Twitter feels a bit like an empty void these days, but it still drums up useful things every now and again. I recently asked for recommendations of “classic” papers in molecular biology. My goal is to add them to the AirTable so that students can track biotechnology progress over time.
Unfortunately, it’s much harder to find “classic” papers than I ever imagined. Cell has a page on “Annotated Classics,” and PNAS has a Classics section that is broken down by field: Genetics, Biochemistry, Medical Sciences, and so on. The best collection is probably Nature’s, but many of their old Classics lists are now outdated or direct to 404 pages.
If you know of any lists on classic biology papers, please let me know!
8 of 31. DNA synthesis is a major bottleneck in building synthetic lifeforms, as I discussed in a recent essay with Travis LaFleur. Oakland-based Ansa Biotechnologies has now made the world’s longest oligonucleotide using enzymes, rather than chemicals. The sequence measures 1,005 bases in length, with a yield of 99.9% at each step. The final pool of oligos contained “approximately 28% sequence-perfect molecules.” Biologists will soon have access to more complex DNA sequences; a boom in foundational technology lifts all boats. GenEng News
9 of 31. Silicon Valley Bank’s collapse is the second largest in U.S. history. In a nutshell:
Deposits in SVB grew 3x between 2019 and 2022, from about $60B to $185B.
The bank used this money to buy bonds. Their portfolio was yielding an average of 1.79%.
The Federal Reserve ramped up interest rates super fast, so the bonds became less valuable. Rates for a 10-year Treasury bond, as of writing, are 3.70%.
Lots of companies needed to withdraw money to meet payroll and to make other payments, but SVB was tied up in bonds, tried to sell them, and then tried to raise $2.3B by selling shares, causing stock sales to crash.
Peter Thiel’s Founders Fund also warned people to withdraw their money from the bank, which “spooked investors,” according to The Guardian.
This is sad news for thousands of startups. Some reports say that 97.3% of the SVB deposits are not FDIC insured (but this number varies across websites). A good write-up on this from Noahpinion: “The longer-term effect of SVB’s collapse on the broader tech sector is likely to be negative, but not catastrophic.”
10 of 31. Lots of gene therapy data this week. A phase 3 study of an AAV5-based therapy for hemophilia B (which causes uncontrolled bleeding) reduced bleeding events by 64 percent; the 54 treated men were monitored between 7 and 18 months post-treatment. The gene therapy was more effective at reducing bleeding events than was the standard of care.
Another phase 3 trial for hemophilia A found that annual bleeding rates “decreased by 84.5% from baseline among the participants” for at least two years after treatment. The study included 112 people who received a single infusion of the gene therapy.
There were also a number of pre-clinical gene therapy studies, including one that used base editing to achieve up to 70% editing efficiencies in organoids carrying a mutation common in patients with cystic fibrosis. And a Phase I study for three patients with late-onset Pompe disease, a rare genetic disorder that causes glycogen to build up inside of cells and causes the heart and skeletal muscles to slowly break down, also looks promising. It showed that a single dose of an AAV8-based therapy could successfully engineer cells to overexpress a gene that breaks down glycogen. The therapy boosted this enzyme’s levels to 235% of baseline after 52 weeks.
11 of 31. AI tools continue to wend their way into science labs. I’m most excited about large language models that accelerate hypothesis generation, or that help scientists find answers to questions faster. This LLM sifts through “dozens of papers” and quickly retrieves answers to scientific questions. 🔻

This tool also looks interesting, but you definitely should not use it. It’s a search engine that scrapes so-called “shadow libraries,” including Sci-Hub, to tread where Google wouldn’t dare. I’m only posting it here so that you know to avoid it.

12 of 31. This is the best explanation I’ve ever seen about why cultivated meat still hasn’t achieved scale, and why the costs might never make sense. Key takeaways:
Amino acids are, surprisingly, a fundamental bottleneck in making cultivated meat. They have to be super pure to avoid contaminating the cells, and “amino acids alone may constitute $7 to $8 per pound of cultivated meat (although there is by no means a consensus about this).”
Getting cells to differentiate into meat, at a large scale, is really difficult. “…going from a 1,000-liter to a 10,000-liter stirred tank reactor involves more vigorous stirring to keep cells and nutrients homogeneously distributed. Cells might not thrive in this more tumultuous environment and could stop growing or break open entirely.”
Short-term funding, and the need for large VC returns, drive a “boom-and-bust” cycle in cultivated meat startups. Lots of knowledge is lost as one company fails and another upstart takes its place with a slightly different approach.
A major challenge is maintaining sterility at scale. “This problem gets worse as scales increase. A bigger batch means a longer, more complex process with more materials — and a higher risk of contamination.”
Recommend. Asterisk Magazine
13 of 31. More papers on AI-based protein design.
The first is a transformer-based protein language model that can identify specific amino acids, within a protein, that have the most “promise” for manipulation. Amino acid changes often render proteins non-functional, or at least less active, so this could be a “promising” tool (heh) to reduce the number of designs that one needs to test. The so-called PROMISE SCORE was tested with both nanobodies and proteins, and it successfully selected site-specific mutagenesis experiments that had a high chance of success.
DeepBIO is a publicly-available tool that integrates “42 state-of-the-art deep-learning algorithms for model training, comparison, optimization and evaluation in a fully automated pipeline.” Lots of these public tools eventually die, sadly, for want of upkeep, time, resources, etc.
Finally, I’m amazed by this recent paper that combined computer simulations and wet-lab experiments to design and make a bacterial protein that can carry out cell division and form spatiotemporal patterns. The scientists plucked out the wildtype gene from E. coli and replaced it with the machine-learning based version, and the cells were perfectly healthy. In other words: They made a synthetic protein that does the same thing as a wildtype protein, but looks nothing like the original. This has huge implications for building truly synthetic cells out of proteins that don’t exist in nature.
Also, David Baker (whose group recently used deep learning to make a new luciferase, or light-emitting, enzyme), gave a talk about his group’s work for the Broad Institute. The video is on YouTube.
14 of 31. Eterna is an online game (there’s also an app) where anyone on the internet can solve puzzles by manipulating RNA sequences and simulating the results. The Eterna team, based at Stanford and Carnegie Mellon, takes the best-performing RNAs and synthesizes them in a lab to see which ones work in real life.
Ribosomes — the big proteins that make other proteins — also, incidentally, use RNA to catalyze peptide bonds. A recent paper used the Eterna platform to crowdsource new RNA sequences that could be used to “improve protein synthesis in vitro and cell growth in vivo,” and many of them actually worked.
15 of 31. The largest private biotech investments from February: A drug delivery company, Aera Therapeutics, raised a $193M Series B. It was founded by Feng Zhang at MIT and aims to use “capsid-like” proteins, naturally encoded within the human genome, to safely transport gene therapies through the body. A biomaterials company, Checkerspot, also raised a $55M Series C. Labiotech
16 of 31. The world’s first genetically-engineered trees have been planted in the U.S. The trees were developed by a startup company, Living Carbon, and are designed to capture and store more atmospheric carbon dioxide than conventional plants. They haven’t been tested in the field yet, so consider their long-term climate impacts with patient, but hopeful, skepticism. The New York Times
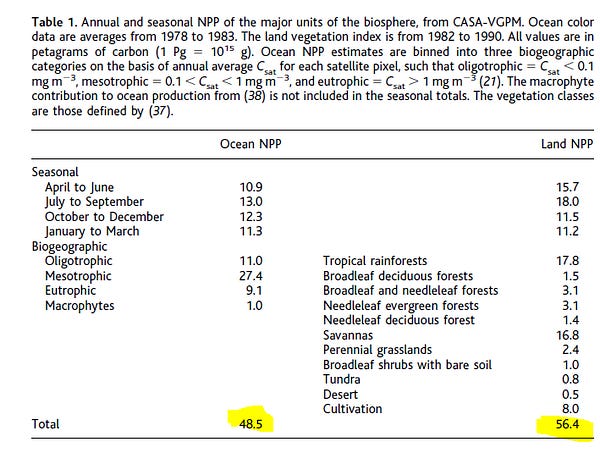
17 of 31. Two types of microbes are needed to make Kombucha: Yeast and Komagataeibacter bacteria (some other bacteria will also do the trick.) The yeast turn sugar into alcohol, while the bacteria eat the alcohol and make cellulose, which looks like a thick, cloudy hockey puck.
Now Komagataeibacter rhaeticus bacteria have been engineered to both produce cellulose and also dye the cellulose a dark black color. The cells were genetically engineered to produce an enzyme, called tyrosinase, that converts L-tyrosine molecules into a eumelanin precursor.

18 of 31. Intellia Therapeutics, the gene-editing company co-founded by Jennifer Doudna, received an ‘OK’ from the U.S. FDA to start a Phase II trial for its gene therapy, called NTLA-2002, to treat an inflammatory disease called hereditary angioedema. The therapy circulates through the bloodstream and is designed to edit a gene in liver cells called KLKB1 to reduce levels of a protein, called kallikrein. A single 25 mg dose of the therapy, tested in three patients, reduced protein levels by 65% after 8 weeks in a prior Phase I/II trial. Reuters
19 of 31. Brain recordings are getting crazy good. In one study, arrays with 4,416 individual electrodes were used to record 3,000 neurons in monkeys at the same time. For context, a neural thread from NeuraLink, in 2019, had more than 3,000 electrodes (on a thread thinner than a human hair) that could record more than 1,000 neurons. The human brain has 86 billion neurons. (Reuters recently reported that the FDA rejected an application from Neuralink to test their device in human subjects.)
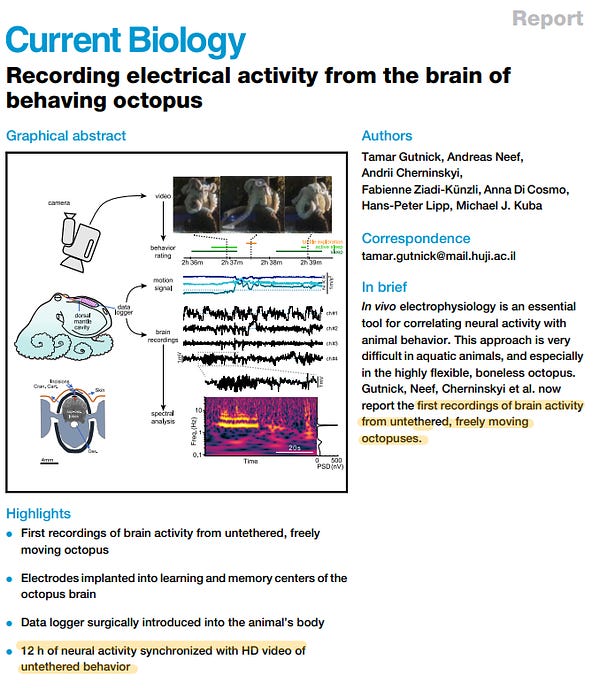
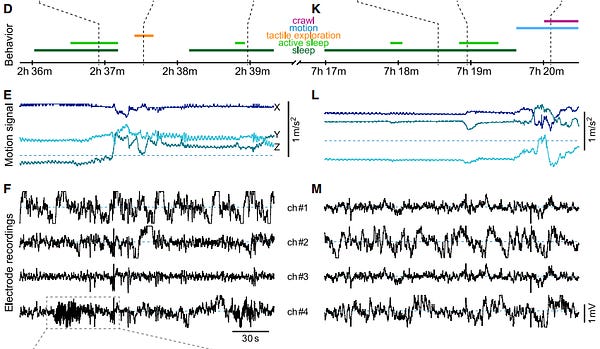
Other recent papers successfully recorded the same neurons from mice over their entire lifespan, which is incredible, and also recorded the brain of a freely-moving octopus for the first time.

20 of 31. As the Earth warms, malaria-causing mosquito ranges have expanded away from the equator by about 2.9 miles per year, on average, over the last century. Gene drives are genetic elements that are transferred to offspring at high rates and can be used to suppress mosquito populations, but I’m not aware of any ongoing field trials for this technology. Burkina Faso, Mali, Ghana, and Uganda are all, apparently, considering experimental releases of mosquitoes that carry gene drives. (Oxitec, a company that riffs on the sterile insect technique to control mosquitoes in Brazil, the Florida Keys, and elsewhere doesn’t use gene drives.) Washington Post
21 of 31. Nearly half of all grant money distributed by the U.S. National Institutes of Health now goes to solicited research aims. The NIH is also funding more clinical trials and more projects over $5M than 20 years ago, suggesting that the discrepancy between the top 1% of projects and “all the rest” has grown over time. Inequalities between old and young investigators, too, are stark (see the NIH Report that I previously edited for New Science). eLife

22 of 31. Caltech biologist, Michael Elowitz, appeared on a podcast with Steven Strogatz for Quanta Magazine. The conversation covers everything from the basics of synthetic biology to questions like: “What does it take to make biology programmable?” Recommend.
Also: I made a mistake in my most recent essay, entitled Biology is a Burrito, and misattributed the source of this title. I later learned that Michael Elowitz has been lecturing on this topic, using this analogy, for at least the last twenty years.
23 of 31. Related to my “Biology is a Burrito” essay: A recent paper calculates how many ATP molecules it takes to build a Trypanosoma brucei parasite, which enters the bloodstream of an unwilling host, eats sugar that floats by, and causes “African sleeping sickness.” In 2010, T. brucei infections killed around 9,000 people, but new cases are now down to the low hundreds annually.
The study’s authors estimate that a single parasite can be “built” from 6x1011 molecules of ATP. A single glucose molecule can make 30 to 32 ATPs, which means that it takes about 19.4 billion sugars to make one parasite.
Of all cellular processes — swimming, eating, digesting — translation (aka making new proteins) is the most energetically expensive. 62% of all energy goes toward making “new biomass,” such as proteins, RNAs, lipids, and so on. Also, 9% of energy is unaccounted for. We don’t know where it’s going. Biology is still a mystery!
24 of 31. Eli Lilly says they’ll cut the price of insulin and will also “cap at $35 a month what patients pay out of their own pockets.”
This is amazing news. In 2016, one-third of people on Medicare in the U.S. had diabetes. Not all take insulin, but Medicare spending on this single compound increased 840% between 2007 and 2017, from $1.4 billion to $13.3 billion per year.
There’s really no reason for this. Insulin should be cheap and plentiful. It was the first human protein made by engineered bacteria. If you’re interested in that story (and the birth of Genentech, and the rise of pretty much all modern biotechnology), I recommend Stephen Hall’s book, Invisible Frontiers.
Basically, in the 1970s, three research groups were all racing to synthesize human insulin in E. coli: Genentech (led by Herbert Boyer at UCSF, together with DNA synthesis maestro Keiichi Itakura and Arthur D. Riggs), William Rutter (also at UCSF) and Walter Gilbert at Harvard.
One of the main reasons why Walter Gilbert lost, and Boyer’s team won, was that the City of Cambridge was afraid of recombinant DNA and banned research that made use of it for months. What began as a 3-month “moratorium” was extended by a city council, unanimously, in September 1976. Federal guidelines at the time also mandated that recombinant DNA work be done in a BSL-4 facility (the highest level, reserved for things like Ebola and Marburg viruses), which is obviously ludicrous in hindsight. The only BSL-4 lab that would work with Gilbert was a British military base in Porton Down, so the whole Harvard team flew there and spent a week or so trying to clone the gene.
How far we’ve come since then! The New York Times

25 of 31. Wine grapes were “probably the first fruit crop domesticated by humans,” about 11,000 years ago. Alcohol is written in our DNA. Washington Post
26 of 31. A company called Retro Biosciences aims “to increase healthy human lifespan by ten years.” A molecule that tunes cellular autophagy (which cells use to break down damaged and abnormal proteins) will enter clinical trials this year, apparently.
The company launched with $180M, all of which came from a single donor: OpenAI’s Sam Altman. Retro is already scooping up some top talent, including José Luis Ricón Fernández de la Puente, aka the blogger, Nintil.
27 of 31. It is possible to make embryos that have three parents: Nuclear DNA from the mother and father, and mitochondrial DNA from a third donor. The goal of this technology, called mitochondrial replacement therapy, is to help couples that have mitochondrial-based genetic diseases produce a healthy child.
The first baby born with three biological parents came in 2016. Making the baby “involved transferring the DNA of a woman’s nucleus into the egg of a donor, which had its own nucleus removed. The baby, a little boy, was born to a woman who carried mitochondrial genes for a disease called Leigh syndrome. Her first two children had died from the disease. But the boy was born healthy.”
At least 19 such embryos have now been transferred into at least 16 women. New evidence suggests that these methods might have unforeseen risks, though, including “reversion,” in which half of a child’s mitochondrial DNA comes from a donor, and the other half, somehow, comes from the mother. MIT Technology Review
Separately: Scientists made egg cells from male mice and successfully fertilized and implanted them into female mice. The results haven’t been published yet, but the work was presented at the Third International Summit on Human Genome Editing in London. Nature
28 of 31. A major challenge in genetic engineering is to predict protein levels, within a cell, from only a DNA sequence. New models can now do this with high accuracy for E. coli bacteria.
But first, some background: Cells use 20 different amino acids to build proteins, and these amino acids are encoded by 64 different codon sequences. CUU, CUC, CUA, UUA, UUG, and CUG in a given RNA sequence all encode leucine. Every organism also has its own ‘preferences’ for codon usage; E. coli encode 47 percent of all leucine amino acids using the CUG codon, while yeast use that codon for just 11 percent of leucines.
A new language model has “learned” these codon usage rules and can predict protein levels from a given sequence. The model was fine-tuned “with over 150,000 functional expression measurements of synonymous coding sequences from three proteins to predict expression in E. coli.” The model was experimentally validated and then used to design gene sequences with optimal expression levels. Amazing.
29 of 31. It’s really difficult to culture most bacteria in the lab. That’s why scientists mainly use yeast — which grows on literally everything — and E. coli. It’s also hard to separate out one specific type of bacteria from a mixed community, like the gut microbiome.
A recent study presents a full-stack solution for the latter. If you take some poop, smear it on a petri dish, and grow a bunch of bacteria, now you can use this “open-source high-throughput robotic strain isolation platform” to sift through all the microbes and pick out individual isolates.
The authors also used a machine learning model to analyze which microbes grow at the same pace as others, or to study which strains only grow when another microbe is nearby. They found many new types of microbial interactions, which might be useful for future microbiome studies.
30 of 31. Kenya is in the midst of its worst drought in four decades, and the country recently approved its first GMO crop. The country banned GMOs and blocked their import back in 2012. The ban lasted until 2019, when the government allowed the import of GM cotton. In October 2022, the government “declared that it would allow farmers to grow pest-resistant GM maize—effectively ending the decade-long ban on GM crops in the country. Since 2015, fall armyworm moths have ravaged maize crops, by one estimate destroying a third of Kenya’s annual production.”
Droughts and extreme flooding will continue to worsen in the next decade. Pests destroy about 40% of all crops, even in the global West. Gene-edited and GMO crops are one solution to the problem. WIRED
31 of 31. A 270-foot long boat, called Falkor, set sail on March 3rd “to look for hydrothermal vents and undiscovered organisms” to understand “how life may have arisen on other planets.” It will sail for forty days. The ship has a remotely operated submarine that can dive to depths of 4,500 meters, or nearly 15,000 feet, and collect samples from the seafloor. New Scientist
You’re a legend! See you soon with a new essay,
— Niko McCarty
(Email | Twitter)
The views expressed in this blog are entirely my own and do not represent the views of any company or institution with which I am affiliated.





Niko, I'm running out of superlatives to describe your newsletter! It just keeps getting better and better...
When are you going to accept levels of subscription? I'm totally opposed to paywalls and monetization of the forum... however, some of us I'm sure would love to support this effort you are making... So long as you keep all the information open to all, don't restrict comment privileges and all the other ills that bedevil the substack environment.
I'm curious as to your level of interest in the philosophy of progress. Or, to put it another way, the assumptions upon which a celebration of progress is built.
In your view, is more knowledge automatically a benefit? Do you see any limit to the amount of knowledge and power human beings can successfully manage?
Are questions like this of interest to you?